MRI: Further selected projects
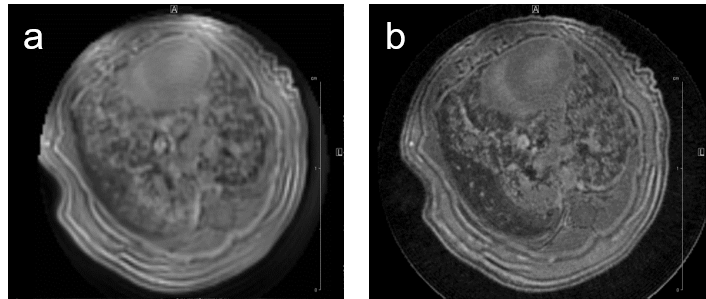
MRI may be used to quantitatively measure lung tumor burden and to follow up e.g. tumor growth. Images of a transgenic mouse model of hyperproliferation in the lung show clearly the replacement of the lung parenchyma Fig. 5.

Fig. 5: Mouse lung MRI. Exemplary axial MR-images of a mouse lung with hyperplasia: (a) respiratory-gated UTE-3D sequence with a repetition time of 8.0 ms, echo time of 20 µs, slice thickness/interslice distance: 0.39/0.39 mm, field of view 2.50x2.70x5.00 cm3 and a matrix of 128/128/128 and (b) respiratory-gated ZTE-3D sequence with a repetition time of 4.0 ms, echo time of 0 µs, slice thickness/interslice distance: 0.16/0.16 mm, field of view 3.00x3.00x4.00cm3 and a matrix of 256/256/256.
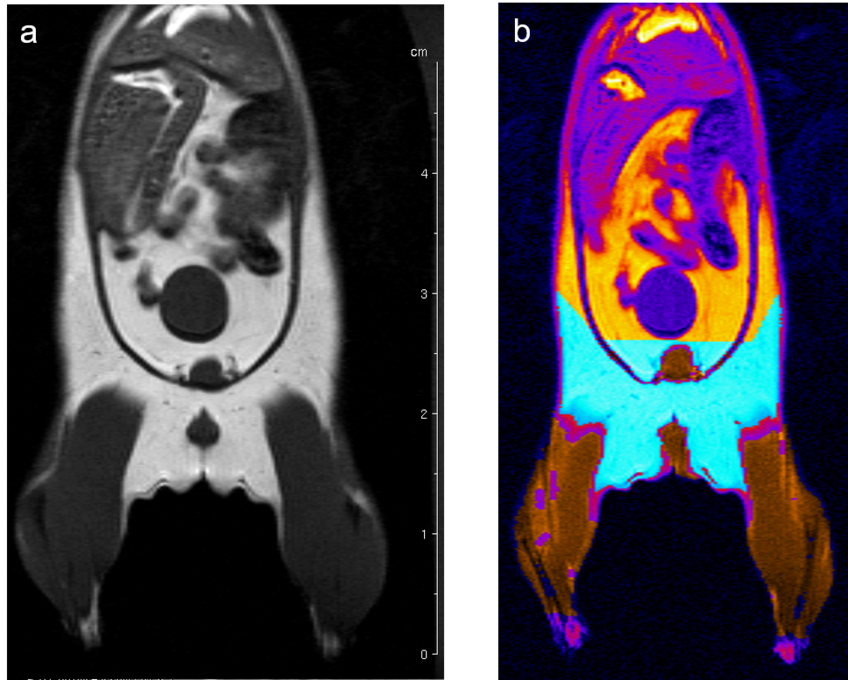
Fat, muscle, tissue-free fluids, and bones generate different signals in response to various radio frequency pulses at distinct magnetic fields, due to their different relaxation properties. We perform MRI studies on mouse using T1-weighted multi-slice-multi-echo (MSME) pulse sequences and analyze the volume of muscle and fat in a well defined and comparable area of the mouse Fig. 6.

Fig. 6: Determination of body fat in mouse. (a) A MSME sequence with a repetition time of 452 ms, echo time of 8.6 ms, field of view 7.0x7.0 cm2 covering the entire body and lower legs, a matrix of 512x256 and slice thickness of 1.0 mm was used to acquire 20 coronal slices with1.0 mm slice thickness and 1.0 mm interslice distance. (b) After evaluation the region in cyan represents the quantified fat and the region in brown the quantified muscle volume.
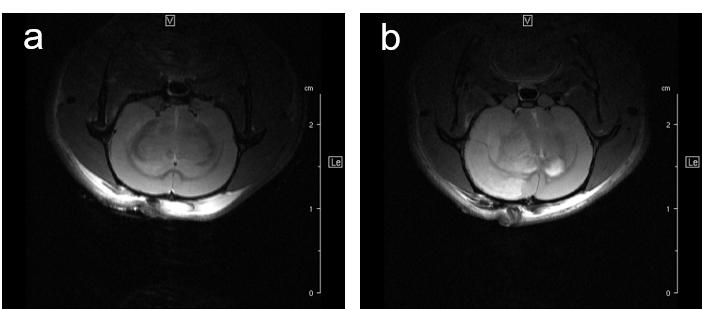
For studying disease pattern of stroke patients, we use a rat animal model with induction of focal cerebral ischemia and follow-up by MRI, Fig.7.

Fig. 7: Infarcted rat brain MRI. For comparison of infarct size of induced infarct right after OP and 24 h later a T2 weighted Spinecho sequence was used: repetition time: 3800 ms, echo time: 18.0 ms, field of view: 35x35 mm2, matrix:512x256 and slice thickness 1.0 mm was used to acquire 12 axial slices.