Molecular Control of Heart and Skeletal Muscle Development
Skeletal muscle forms the physical basis for all coordinated locomotion, a key feature of all higher animals. By weight, skeletal muscle is the dominant organ system of animals and contributes importantly not only to locomotion, but also to energy metabolism. Understanding the development of skeletal muscle into distinct lineages, some of which retain self-renewing properties allowing muscle repair, their functional adaptation into fast and slow fibres, and the tightly interlinked regulation of muscle protein homeostasis with physical demand and global energy demand all greatly contribute to a fundamental new insight.
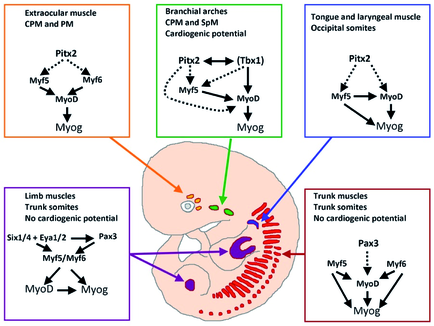
Skeletal muscle cells of higher vertebrates arise from three different locations: segmented somatic paraxial mesoderm, unsegmented paraxial head mesoderm, and prechordal mesoderm. The skeletal muscles of the trunk and limbs are derived from the segmented somitic paraxial mesoderm, the somites, which form on either side of the neural tube in vertebrate embryos. Upon further development, somites differentiate into a dorsal part, called the dermomyotome, which retains an epithelial structure and a ventral part, called the sclerotome, which breaks up into mesenchyme that contributes to the axial skeleton of the embryo. The first muscle cells are formed in the myotome, which is located directly underneath the dermomyotome by cells that delaminate from the dermomyotome.
Determination and terminal differentiation of all skeletal muscle cells are governed by a network of four myogenic regulatory factors (MRFs: Myf5, Mrf4, Myf6, MyoD, and myogenin). Myf5 and MyoD are generally believed to act as determination genes, while myogenin is essential for the terminal differentiation of committed myoblasts. Mrf4/Myf6 seems to play a dual role. It has long been considered to be a differentiation gene, but it is also expressed in undifferentiated cells in which it might act as a determination gene.
A number of transcription factors have been identified, which are located upstream of MRFs. None of these factors is located exclusively in the progenitor cell population that will give rise to muscle cells, which suggests that these transcription factors prepare the stage for additional actors (such as MRFs) to initiate myogenesis or act together with additional transcription factors in a combinatorial manner to allow activation of MRFs. In both cases instructive or permissive inductional cues are necessary to achieve stable self-sustained activation of MRFs and muscle cell formation.

Heart formation encompasses an orchestrated series of molecular and cellular events, and thus even subtle alterations in this process can lead to serious cardiac disorders. Defects in very early stages of cardiac development often result in a failure of the cardiovascular system that leads to embryonic lethality whereas alterations during later stages are generally associated with congenital heart defects seen in embryos, which survive to birth. Congenital heart defects are the most common of all birth defects in humans. Adult cardiac muscle lacks sufficient regenerative potential to cope with acute, massive loss of cardiomyocytes that follow myocardial infarction, which results in formation of non-functional scar tissue. Furthermore, progressive loss of cardiomyocytes during aging and chronic disease could lead to heart failure, a major cause of hospitalization and death in elderly people.
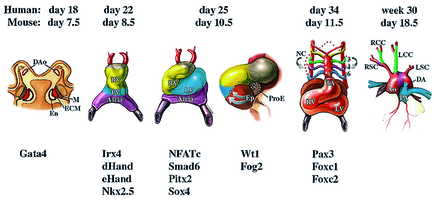
Heart tissue is mostly derived from mesodermal cells that migrate anterior-laterally from the primitive streak during early gastrulation. The anterior lateral mesoderm splits into somatic and splanchnic layers; the splanchnic mesoderm forms the cardiac crescent that later gives rise to the heart myocardium. Ventral folding of the embryo and growth of the cardiac epithelium generate the tubular heart, with separate inflow (venous) and outflow (arterial) poles. In all vertebrates, the linear heart tube undergoes rightward looping, during which the heart tube grows dramatically in length. Following looping, the heart undergoes a transition from a simple tubular pump into a multi-chambered structure via substantial growth and remodeling events. The vertebrate heart is formed from two distinct mesoderm populations or “heart fields”, which arise from a common origin, and express both distinct and overlapping molecular markers. The earliest population of cardiac progenitors, the first heart field, corresponds to the anterior lateral mesoderm and, subsequently, to the cardiac crescent. Ultimately, cells from the first heart field contribute to the left ventricle and atria. An additional source of cardiac precursors, the second (or secondary/anterior) heart field derives from mesoderm located medial to the cardiac crescent. These cells contribute primarily to the cardiac outflow tract, right ventricle and atria.
Heart development is governed by a core set of evolutionarily conserved transcription factors (NK2, MEF2, GATA, Tbx, and Hand) that direct cardiac cell fates, the expression of genes encoding sarcomeric proteins, and the morphogenesis of cardiac structures (Olson, 2006). The core set of transcription factors also regulate each other’s expression, thereby stabilizing and reinforcing the cardiac gene program.
Dozens of other transcription factors contribute to cardiogenesis by serving as accessory factors for core regulators or by fulfilling yet unknown functions (McFadden, 2002). Morphogenetic processes need to be coordinated with processes that direct formation of the primary heart tube and the differentiation of cardiomyocyte differentiation. Surprisingly, the known core set of cardiac transcription factors mainly controls the expansion of chamber myocardium but not the initial differentiation of cardiomyocytes and formation of the primary heart tube. Although expression analysis of early heart development has yielded lists of potential candidate genes that might control this primary process, a clear functional role of these genes for cardiomyocyte differentiation has not been established.
Cardiac progenitor cells respond to positive signals from surrounding tissues such as the anterior endoderm. It has been shown that bone morphogenic protein (BMP) and fibroblast growth factor (FGF) signals secreted by the anterior endoderm are required for cardiac differentiation in the splanchnic mesoderm.
In contrast, studies in chick and xenopus embryos had demonstrated that the canonical Wnt signaling pathway plays an important role in suppressing cardiac mesoderm differentiation. Our current notion describes a multiphasic role for Wnt signaling in the induction of mesoderm formation and expansion of the cardiovascular progenitor pool along with inhibition
of myocardial differentiation. Bmp, Fgf and Wnt signaling pathways are clearly the most intensively studied evolutionary conserved signaling mechanisms that regulate cardiogenesis in both vertebrates and invertebrates. However, the precise molecular mechanisms and the crosstalk between these signaling pathways are not entirely clear.

Research projects in the department aim to identify new molecular circuits that control heart and skeletal muscle development at all levels, on which genetic information is realized. Previous research efforts have mostly focused on the role of transcription factors such as MRFs. In order to place these efforts into a larger perspective we now put more emphasis on the regulation of the chromatin structure, which is instrumental for the activation of the myogenic program, and on posttranscriptional control events regulating mRNA stability and translation via miRNAs and RNA binding proteins.